Standard JOB submission
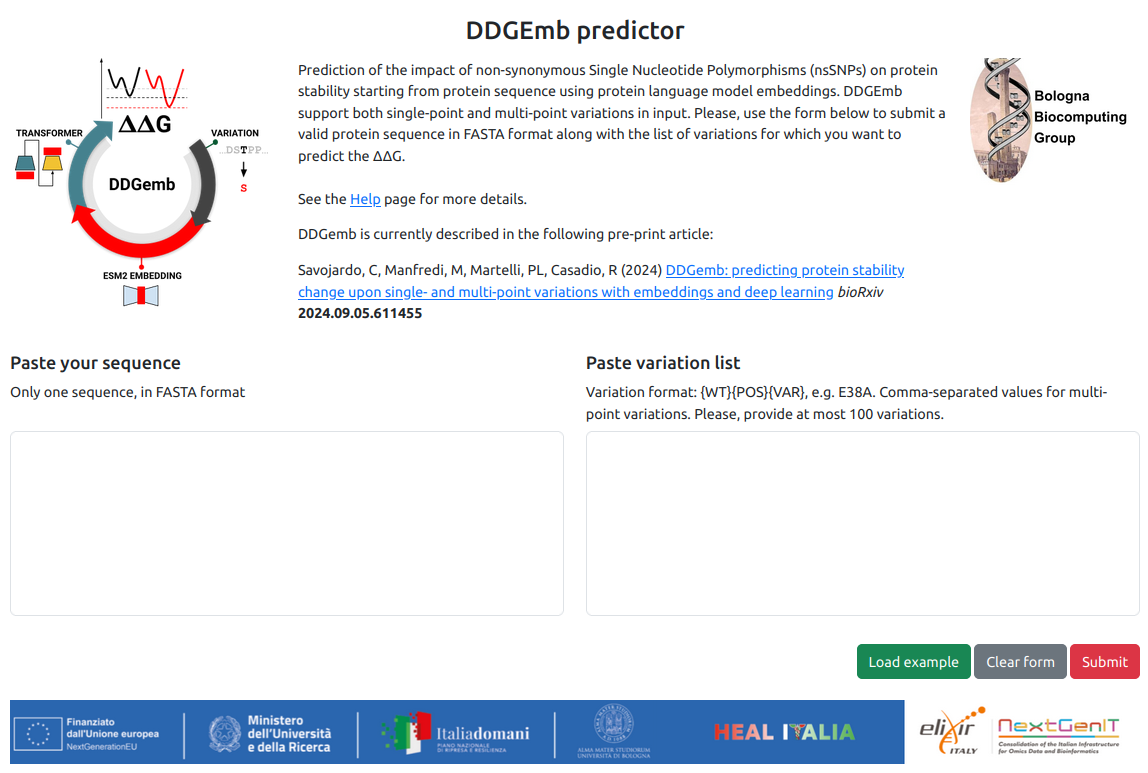
From the DDGemb home page, a standard DDGemb job including up to 100 variations on a single protein sequence can submited.
JOB submission requires:
- A protein sequence in FASTA format
- A list of variations, one per line
Variations must be provided in the {WT}{POS}{VAR} format, e.g. S11C. Multi-point variations are reported in the same line, comma-seprated, e.g. S11C,F18L. Variant positions are specified starting from 1 to the length of the sequence.
Upon submission, data will be checked for validity. In particular, the following validation checks are performed:
- The protein sequence must be shorter than 2500 residues.
- No more than 100 variations are provided.
- Variant positions are not out of the range of the sequence.
- Individual variations in multi-point variants affect different sequence positions.
- For all variant positions, wild-type residues are consistent with those in the sequence.
- All variant residues are in the set of standard amino acids and different from the respective wild-type.
Once validated, the input sequence and variations are provided to DDGemb for ΔΔG prediction. After submission, a unique identifier is assigned to the JOB and can be then used to retrieve JOB results later.
Example data can be loaded using the "Load example" button.
Visualization of JOB results
Once completed, the JOB results will be automtically visualized.
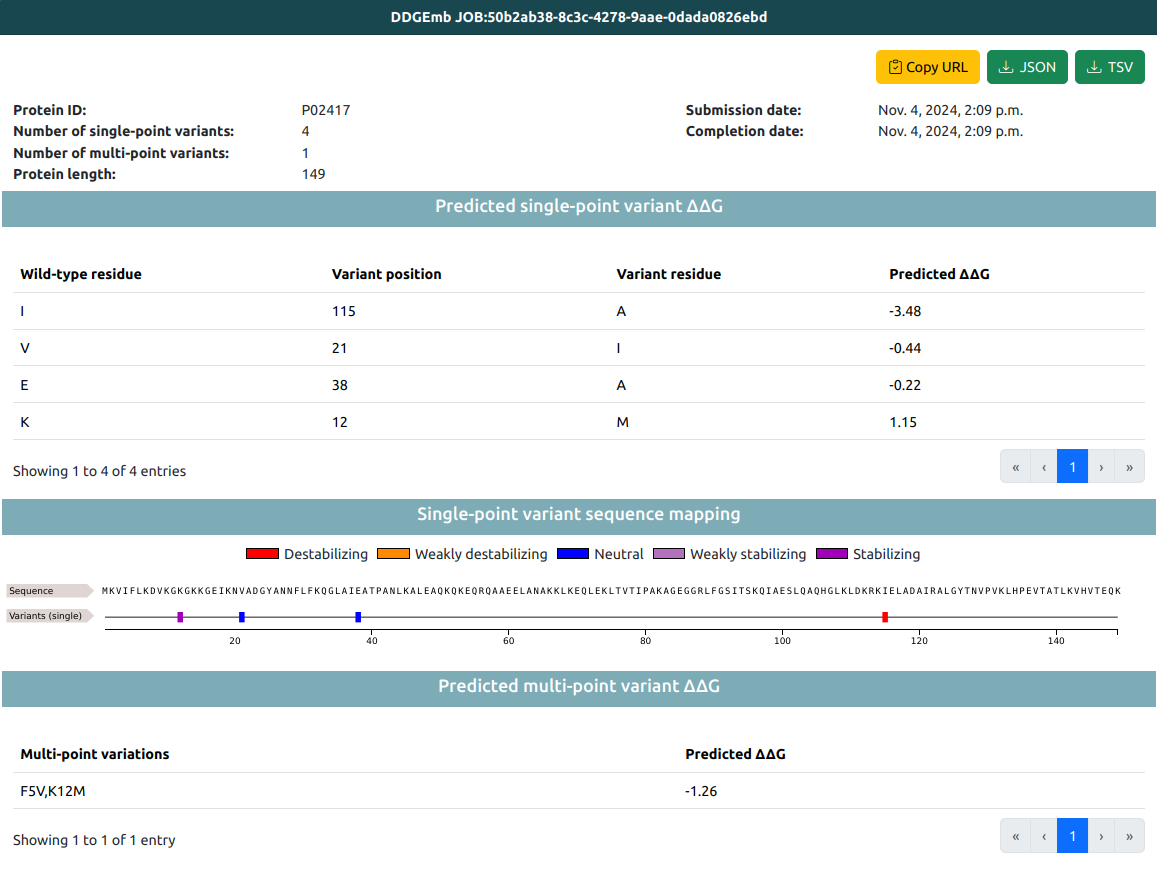
A typical result page contains the following information:
- General information about the submitted JOB.
- Results of ΔΔG prediction of single- and multi-point variations.
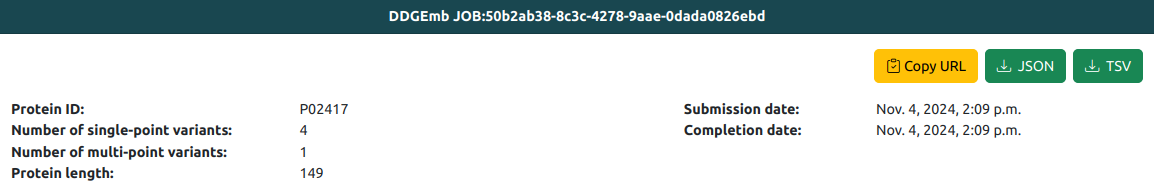
JOB general info tab reports:
- The protein identifier, as reported in the input FASTA file.
- The number of single-point variations processed by the JOB.
- The number of multi-point variations processed by the JOB.
- The protein length.
- The JOB submimission and completion time
On top of panel, three buttons are available for
- Copy the URL of the JOB result page, for future retrieval.
- Download the results in JSON and TSV formats.
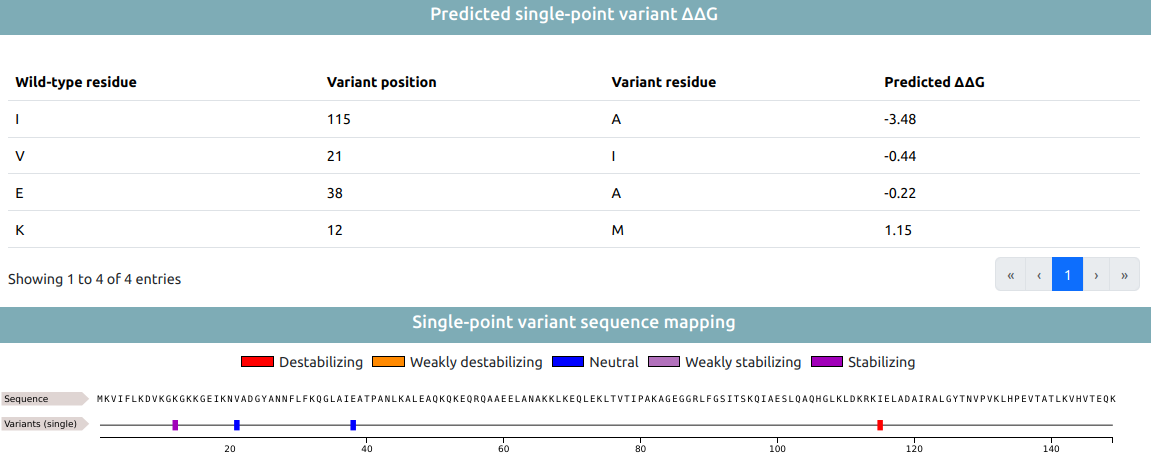
Result of ΔΔG prediction is then reported for single-point variations included in the JOB (if any). Theese are reported in two ways: tabular format and visualized along the sequence (using a FeatureViewer).
Tabular data include, for each single-point variations:
- The wild-type residue.
- The variant position.
- The variant residue.
- The predicted ΔΔG.
Sequence visualization based on FeatureViwer reports the different variants along the sequence as features colored according to the value of the predicted ΔΔG. Five classes are identified:
- Destabilizing variations (red), when ΔΔG < -1
- Weakly destabilizing variations (orange), when -1 ≤ ΔΔG < -0.5
- Neutral variations (blu), when -0.5 ≤ ΔΔG ≤ 0.5
- Weakly stabilizing variations (pink), when 0.5 < ΔΔG ≤ 1
- Stabilizing variations (purple), when ΔΔG > 1
Result of ΔΔG prediction is then reported for multi-point variations included in the JOB (if any). Theese are reported in tabular format, including:
- The multi-point variation (i.e., a comma-separated list of single-point variations).
- The predicted ΔΔG.
Batch JOB submission
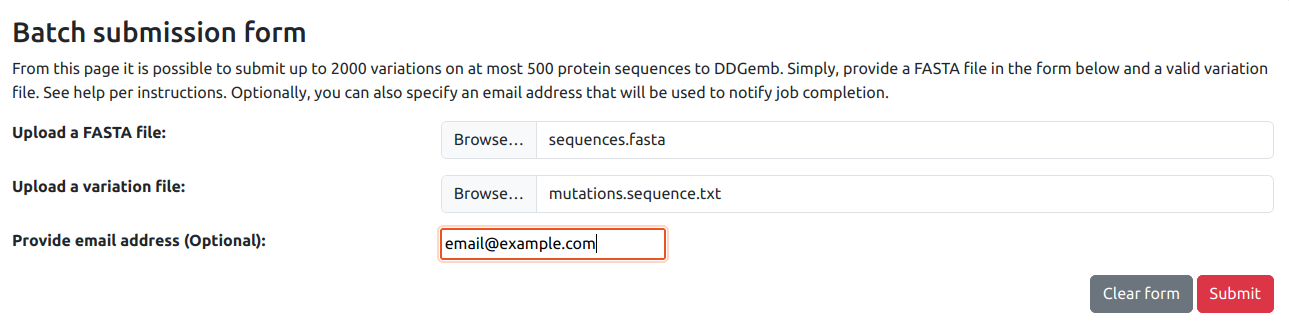
Using the DDGemb batch submission form it is possible to analyze up to 2000 variations (either single- or multi-point) occurring on at most 500 protein sequences.
The input is provided by means of two different files:
- A FASTA file, containing the set of proteins sequences.
- A variation file, containing the list of variations to be analyzed.
The provided input FASTA file must contain uniq protein sequences. Duplicated sequences (even using a different IDs) are not allowed.
Variations are reported in the same {WT}{POS}{VAR} (e.g., V21I) format as in the single-sequence case, with one additional column reporting the identifier of the protein, as included in the corresponding FASTA file, for example:
P02417 I115A
P02417 V21I
P02417 E38A
P02417 K12M
P02417 F5V,K12M
P03069 E270Q
P03069 K276A
and FASTA file will be:
>P02417
MKVIFLKDVKGKGKKGEIKNVADGYANNFLFKQGLAIEATPANLKALEAQKQKEQRQAAE
ELANAKKLKEQLEKLTVTIPAKAGEGGRLFGSITSKQIAESLQAQHGLKLDKRKIELADA
IRALGYTNVPVKLHPEVTATLKVHVTEQK
>P03069
MSEYQPSLFALNPMGFSPLDGSKSTNENVSASTSTAKPMVGQLIFDKFIKTEEDPIIKQD
TPSNLDFDFALPQTATAPDAKTVLPIPELDDAVVESFFSSSTDSTPMFEYENLEDNSKEW
TSLFDNDIPVTTDDVSLADKAIESTEEVSLVPSNLEVSTTSFLPTPVLEDAKLTQTRKVK
KPNSVVKKSHHVGKDDESRLDHLGVVAYNRKQRSIPLSPIVPESSDPAALKRARNTEAAR
RSRARKLQRMKQLEDKVEELLSKNYHLENEVARLKKLVGER
Before submission, the user is asked to optionally provide an e-mail address to receive a notification of JOB completion.
Once completed, JOB results can be downloaded in JSON and TSV formats.
Search for previously submitted JOBs
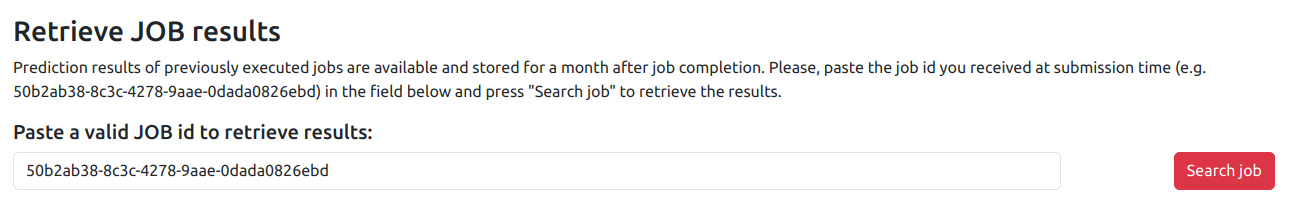
Results of previously submitted and completed JOBs can be retrieved using the JOB search page.
To retrieve a JOB (either interactive or batch), you must provide the unique JOB id obtained at JOB submission, e.g. 50b2ab38-8c3c-4278-9aae-0dada0826ebd. If the JOB id is valid, you will be automatically redirected to the JOB result page.